The Loraine Lab at UNC Charlotte has trained and supported more than 50 members since 2008. Some of them are listed below.
 Dylan Marrotte graduated from UNC Charlotte with a Bachelor’s degree in Biology in 2023. During his time as an undergraduate, he conducted research with bats and investigated the developmental biology of the Phyllostomidae family, exploring brain development during gestation across different species using advanced imaging techniques.
Dylan Marrotte graduated from UNC Charlotte with a Bachelor’s degree in Biology in 2023. During his time as an undergraduate, he conducted research with bats and investigated the developmental biology of the Phyllostomidae family, exploring brain development during gestation across different species using advanced imaging techniques.
Dylan joined the Loraine Lab in 2024 during his first year as a master’s student. In his role of research assistant, he produced outreach materials, assisted with data management, and contributed to software development and testing.
Dylan is now focused on finishing the last year of his Bioinformatics master’s program here at UNC Charlotte.
 Molly Davis received her Bachelor’s degree, majoring in Biology and minoring in Bioinformatics, from UNC-Charlotte in 2021. During her time as an undergraduate, she entered in the Bioinformatics Technology Certificate Program as an Early Entry Graduate which led to the completion of her Master’s degree in Bioinformatics in 2022.
Molly Davis received her Bachelor’s degree, majoring in Biology and minoring in Bioinformatics, from UNC-Charlotte in 2021. During her time as an undergraduate, she entered in the Bioinformatics Technology Certificate Program as an Early Entry Graduate which led to the completion of her Master’s degree in Bioinformatics in 2022.
Molly’s first experience with the Loraine Lab was as a Bioinformatics Analyst intern under the guidance of Dr. Robert Reid. She researched heat stress in different tomato species and analyzed sequencing data. This internship turned into a Bioinformatics Programmer position after her graduation where she worked on a Nextflow pipeline to then analyze alternative splicing locations in IGB based on different stress conditions.
Molly is now pursuing a Biostatistics Masters Certification at the University of Louisville.
 Kaushik Gopu received his Bachelor of Technology degree in Computer Science from GITAM Deemed University in India. He then joined the lab in 2023 as a master’s student in Computer Science at UNC Charlotte. During his time with our lab, he was instrumental in developing a new file parser that can process CRAM file format in the Integrated Genome Browser. He also worked on adding new features to the Integrated Genome Browser and updating an IGB App called ProtAnnot. Additionally, he contributed to improving our use of OSGi frameworks by trying out and reviewing various online tutorials discussing declarative services annotations and their use.
Kaushik Gopu received his Bachelor of Technology degree in Computer Science from GITAM Deemed University in India. He then joined the lab in 2023 as a master’s student in Computer Science at UNC Charlotte. During his time with our lab, he was instrumental in developing a new file parser that can process CRAM file format in the Integrated Genome Browser. He also worked on adding new features to the Integrated Genome Browser and updating an IGB App called ProtAnnot. Additionally, he contributed to improving our use of OSGi frameworks by trying out and reviewing various online tutorials discussing declarative services annotations and their use.
Kaushik’s first job following his graduation from UNC-Charlotte is software engineer for VCO Systems in Roswell, Georgia.
Logan Weidenhammer joined the group in May 2020. In the following months, they upgraded IGB testing and user documentation, helped redesign the IGB App Store interface to use a new faceted search interface, and assessed usability designs for many other projects.
In May, 2020, Logan completed their Graduate Certificate in Human-Computer Interaction
from the Software and Information Systems Department at UNC Charlotte.
As an undergraduate, Logan studied psychology and did post-bac coursework in biology and health sciences.
 Philip Badzuh joined the lab as a Research Assistant in 2017. He created computational pipelines to automate data collection, transformation, and analysis. He also cultivated antibiotic-resistant Burkholderia strains and measured their growth rates in the lab to assist with Cystic Fibrosis research. He graduated with his Master’s degree in Bioinformatics in 2022.
Philip Badzuh joined the lab as a Research Assistant in 2017. He created computational pipelines to automate data collection, transformation, and analysis. He also cultivated antibiotic-resistant Burkholderia strains and measured their growth rates in the lab to assist with Cystic Fibrosis research. He graduated with his Master’s degree in Bioinformatics in 2022.
Philip’s first job after graduating is Full Stack Software Engineer at AmeriSave Mortgage Corporation in Charlotte.
 Omkar Marne joined in April 2021, about a year after earning his MS in Computer Science at UNC Charlotte. In just a few months, he contributed to three major projects in the group. Working in client-side java, he helped improve the Integrated Genome Browser user interface. Working in server-side python, using the Django framework, he improved the IGB App Store and co-developed the first version of the new Track Hub Facade site.
Omkar Marne joined in April 2021, about a year after earning his MS in Computer Science at UNC Charlotte. In just a few months, he contributed to three major projects in the group. Working in client-side java, he helped improve the Integrated Genome Browser user interface. Working in server-side python, using the Django framework, he improved the IGB App Store and co-developed the first version of the new Track Hub Facade site.
Omkar is currently pursuing his Ph.D. in Bioinformatics at UNC-Charlotte.
 Irvin Naylor joined the IGB team in August 2020 during his senior year of undergraduate studies in computer science at UNC Charlotte, and continued in the lab part-time following graduation. During his tenure, Irwin contributed to several projects. On the IGB project, he improved our testing documentation, added new genomic data to IGB, and helped test and prepare the IGB 9.1.8 release. He also helped maintain the IGB build system, updating the POM build configuration files. He created a new IGB demo App to demonstrate the filtering features of the API. And in his final months in the lab, he worked with a team of two others to develop an entirely new application that lets IGB users visualize Track Hubs, a Quickload-like mechanism for sharing genomic data on-line via HTTP.
Irvin Naylor joined the IGB team in August 2020 during his senior year of undergraduate studies in computer science at UNC Charlotte, and continued in the lab part-time following graduation. During his tenure, Irwin contributed to several projects. On the IGB project, he improved our testing documentation, added new genomic data to IGB, and helped test and prepare the IGB 9.1.8 release. He also helped maintain the IGB build system, updating the POM build configuration files. He created a new IGB demo App to demonstrate the filtering features of the API. And in his final months in the lab, he worked with a team of two others to develop an entirely new application that lets IGB users visualize Track Hubs, a Quickload-like mechanism for sharing genomic data on-line via HTTP.
Irvin now works as an ITS Application Developer at Selective Insurance.
 Chester Dias joined Loraine Labs during the spring semester of 2020. His first project focused on automating deployment and updates for the IGB App Store. As part of this project, he developed ansible playbooks for deploying resources via Amazon Web Services and updating deployed code. He then applied knowledge gained from this project to automate deployment of the BioViz.org Web site and many others.
Chester Dias joined Loraine Labs during the spring semester of 2020. His first project focused on automating deployment and updates for the IGB App Store. As part of this project, he developed ansible playbooks for deploying resources via Amazon Web Services and updating deployed code. He then applied knowledge gained from this project to automate deployment of the BioViz.org Web site and many others.
After graduating in May of 2021, Chester joined Deloitte as a Solutions Specialist.
 Chirag Shetty joined the Loraine Lab shortly after graduating with an MS in Computer Science from University of Alabama, Birmingham in May 2020. During the next eleven months, he contributed to many projects. To start, he took over developing the IGB App Store, finalizing several new features of the software and helping to release version 2.0, which introduced major changes. In his final months, he co-developed a new application that allows IGB users to view data from UCSC Track Hubs, a Quickload-like mechanism for sharing genomic data on-line via HTTP.
Chirag Shetty joined the Loraine Lab shortly after graduating with an MS in Computer Science from University of Alabama, Birmingham in May 2020. During the next eleven months, he contributed to many projects. To start, he took over developing the IGB App Store, finalizing several new features of the software and helping to release version 2.0, which introduced major changes. In his final months, he co-developed a new application that allows IGB users to view data from UCSC Track Hubs, a Quickload-like mechanism for sharing genomic data on-line via HTTP.
Chirag now works as a Data Engineer for Sense, based in Boston, Massachusetts.
 Sai Supreeth Segu joined the lab in October 2020, after graduating from UNC Charlotte with an MS degree in Computer Science. He worked with us on a part-time basis until March 2021, when he obtained full-time employment as a software engineer. During this time, he contributed to the App Store project and helped improve the GenoViz SDK build system and IGB code bases.
Sai Supreeth Segu joined the lab in October 2020, after graduating from UNC Charlotte with an MS degree in Computer Science. He worked with us on a part-time basis until March 2021, when he obtained full-time employment as a software engineer. During this time, he contributed to the App Store project and helped improve the GenoViz SDK build system and IGB code bases.
Supreeth is currently working as an Associate at Goldman Sachs, based in Salt Lake City, Utah.
 Chaitanya Kintali joined the group in May 2019 following his first semester at UNC Charlotte.
Chaitanya Kintali joined the group in May 2019 following his first semester at UNC Charlotte.
During his tenure in the Loraine Lab, Chaitanya primarily worked on BioViz Connect, performing a lot of the back end engineering. For example, he designed and implemented BioViz Connect’s mechanism of updating its list of IGB-related Apps deployed on CyVerse.
Since earning his MS in Computer Science, Chaitanya has worked as a Software Engineer at Amazon as well as a Senior Software Engineer at Adtechcorp.
 Sameer Shanbhag joined the lab in May 2019 following his first semester at UNC Charlotte, where he is pursuing a masters degree in Computer Science. Sameer received his undergraduate degree in Engineering from the SIES Graduate School of Technology in India.
Sameer Shanbhag joined the lab in May 2019 following his first semester at UNC Charlotte, where he is pursuing a masters degree in Computer Science. Sameer received his undergraduate degree in Engineering from the SIES Graduate School of Technology in India.
Prior to starting at UNC Charlotte, Sameer worked at at Xoriant Solutions as a Software Engineer. At Xoriant, he mainly worked as a consultant for client site Morgan Stanley developing middleware service to provide business and financial information for the company.
In the Loraine Lab, Sameer played a key role in developing the App Store for IGB and the Genome Dashboard software.
Sameer now works as a Senior Software Engineer at Walmart Global Tech, based in Sunnyvale, California.
 Noor Zahara joined the group during the summer of 2019 and worked with us until she graduated in Dec 2020. During summer of 2020, she did an internship at Amazon.
Noor Zahara joined the group during the summer of 2019 and worked with us until she graduated in Dec 2020. During summer of 2020, she did an internship at Amazon.
Noor made many improvements to the IGB software, and co-developed the IGB App Store. She also helped with releases and testing, and helping new joiners get familiar with code and tools used in the lab.
After graduation, Noor re-joined Amazon as a software engineer.
 Jay Chamma joined the lab as a summer intern in May 2020 while working toward his undergraduate degree in Computer Science from UNC Charlotte. Prior to UNC Charlotte, he earned an Associate in Science (A.S.) degree from Central Piedmont Community College in Charlotte.
Jay Chamma joined the lab as a summer intern in May 2020 while working toward his undergraduate degree in Computer Science from UNC Charlotte. Prior to UNC Charlotte, he earned an Associate in Science (A.S.) degree from Central Piedmont Community College in Charlotte.
During his internship, Jay made videos for the IGB YouTube channel and helped identify and fix problems with IGB on the Windows operating system. He also helped identify research papers where IGB played a role.
In addition to CS, Jay is deeply interested in genomics and applications of genomics to health care. Jay now works as a Full Stack Application Developer for U.S. Bank.
 Pooja Nikhare joined the group in fall of 2019, her first semester at UNC Charlotte, where she is pursuing her masters degree in Computer Science.
Pooja Nikhare joined the group in fall of 2019, her first semester at UNC Charlotte, where she is pursuing her masters degree in Computer Science.
Prior to coming to Charlotte, Pooja worked for several years at the Centre for Development of Telematics in India, where she rose to the position of Senior Research Engineer. She worked mainly on networking related projects, such as developing tools and algorithms for reducing network faults and understandability.
Pooja received her undergraduate degree in Computer Science and Engineering from National Institute of Technology, Tiruchirappalli.
On the IGB project, Pooja is used her advanced programming skills to develop and maintain IGB Apps. She has also played a vital role in testing and validation of the IGB App Store.
Pooja now works as a Software Engineer at Microsoft.
 Prutha Kulkarni joined the IGB development team in May 2019 following her first semester at UNC Charlotte in the Computer Science program.
Prutha Kulkarni joined the IGB development team in May 2019 following her first semester at UNC Charlotte in the Computer Science program.
Prutha received her Bachelor of Engineering in Information Technology from Pune University in 2015. Following this, she worked as a Software Developer and Technical Consultant at FinIQ, also in Pune. At FinIQ, she developed resources such as an in-house syntax highlighter (.Net) and graphical visualization methods using D3, HTML, and CSS for better understanding of products and company data.
In the Loraine Lab, Prutha developed and maintained our collection of IGB Apps and played a key role in improving our Bitbucket pipelines-based build process. She received her MS degree in Computer Science in May 2020.
Prutha now works as a Senior Full Stack Engineer at GPS DataViz.
 Pawan Bole joined the group in fall 2018, near the end of his first semester at UNC Charlotte.
Pawan Bole joined the group in fall 2018, near the end of his first semester at UNC Charlotte.
Before coming to UNC Charlotte, Pawan worked as a Software Developer for Tavisca Solutions Pvt. Ltd in Pune, India. At Tavisca, Pawan contributed to multiple projects, mainly focusing on developing Web applications using .Net.
Pawan earned his undergraduate degrees in Computer Science from University of Mumbai, and then earned an Advanced Computing credential from the Center for Development of Advance Computing (CDAC) in Mumbai.
Pawan worked on the BioViz Connect project and also contributed to the IGB code base. He received his MS degree in Information Technology in May 2020.
Pawan now works as a Software Engineer at Meta.
 Srishti Tiwari joined the lab in August 2018 during her first semester at UNC Charlotte as a graduate student in the Information Technology Masters program.
Srishti Tiwari joined the lab in August 2018 during her first semester at UNC Charlotte as a graduate student in the Information Technology Masters program.
Srishti received her undergraduate degree in Electronics (B.E.) from Mumbai University. In addition, she holds a PG-Diploma in Advanced Computing from CDAC-ACTS. During her final year project, she worked on a DSP processor for processing analog speech. Following graduation, she worked for several years as a Senior Software Engineer at such companies as MITS Global, BNP Paribas ISPL (on contract with MITS Global), HERE Solutions and SmartStream Technologies.
During her tenure in the lab, Srishti contributed to many projects, including IGB App Store, BioViz Connect, and the IGB code base itself. She received her MS degree in Information Technology in May 2020.
Srishti now works as a Software Engineer at Salesforce.
 Shamika Kulkarni joined the lab in May 2019 following her first semester as a graduate student at UNC Charlotte.
Shamika Kulkarni joined the lab in May 2019 following her first semester as a graduate student at UNC Charlotte.
Shamika earned her Bachelor of Engineering in Electronics and Telecommunication from S.I.E.S. Graduate School of Engineering, University of Mumbai, in May 2016.
Following this, she worked at TATA as a Consultancy Assistant System Engineer in for the State Bank of India. In this role, she focused on developing and handling software for banking transactions.
In the Loraine Lab, Shamika developed and improved several IGB Apps. She played a key role in developing demo Apps illustrating aspects of IGB’s internal Java API. She also took the lead on testing as we rolled out new products and projects.
Shamika now works as an Associate at Goldman Sachs.
 Narendra Kumar Vankayala joined the group during the spring semester of 2019. He earned his masters degree in computer science in December 2019.
Narendra Kumar Vankayala joined the group during the spring semester of 2019. He earned his masters degree in computer science in December 2019.
Following graduation, he joined Capital Group as a software engineer, focusing on solving engineering problems related to machine learning and artificial intelligence.
Prior to starting his masters, he worked as a Software Engineer at Mindtree in Bengaluru, India. Narendra now works as a Data Product Manager at Capital Group.
 Sai Charan Reddy Vallapureddy joined the IGB team during during spring, summer, and fall of 2019. He received his masters degree in Computer Science in Dec 2019.
Sai Charan Reddy Vallapureddy joined the IGB team during during spring, summer, and fall of 2019. He received his masters degree in Computer Science in Dec 2019.
Charan worked on many different projects, including: migrating a gene lookup REST service to use AWS, developing new code for the IGB App Store, and implementing a new BAM index visualization for Integrated Genome Browser.
Charan now works as a Software Developer for Insurity, based in Austin, Texas.
 Riddhi Jagdish Patil joined the Integrated Genome Browser development group during spring and fall of 2019. She received her MS in Computer Science in December 2019.
Riddhi Jagdish Patil joined the Integrated Genome Browser development group during spring and fall of 2019. She received her MS in Computer Science in December 2019.
Riddhi contributed to several aspects of the IGB code base. She co-wrote a new IGB endpoint for installing IGB Apps from IGB App Store. She also co-wrote or improved several aspects of the IGB App Store itself.
Following graduation, Riddhi moved to Seattle for a new position as a software engineer at Amazon.
 Srishtee Marotkar joined the Loraine Lab software development group in fall of 2018 during her first semester at UNC Charlotte as a graduate student in Computer Science.
Srishtee Marotkar joined the Loraine Lab software development group in fall of 2018 during her first semester at UNC Charlotte as a graduate student in Computer Science.
Srishtee earned a Bachelor of Technology degree in Electronics Engineering from the University of Pune. Following this, she joined Cognizant Technology Solutions as a Software Engineer.
At Cognizant, she developed Web applications for company clients, mainly in finance and banking industries.
Srishtee is graduating in fall of 2019. After that, she plans to join Amazon as a full-time developer, based in Seattle.
Jill Jenkins joined the lab in October 2019, shortly after we expanded the computational lab into new space in the Core Lab building at Kannapolis.
Jill is pursuing a masters degree in Bioinformatics. In addition to bioinformatics, Jill brings a deep knowledge of biology and how to retrieve biological data from on-line database, notably Ensembl and BioMart.
In addition to pitching in on testing for the IGB project, Jill is primarily serving as Bioinformatics Data Manager, performing the complex tax of searching out new data sets to include the IGB Quickload data sharing system. She also writes code (python) to manage and process data.
Jill now works as a Senior Data Scientist for Premier Inc.
Kiran Korey joined the Integrated Genome Browser team in April of 2018 during his first semester as a Computer Science graduate student at UNC Charlotte.
Kiran earned a Bachelor of Information Science in 2015 from B V Bhoomaraddi College of Engineering and Technology, where he studying operating systems, software engineering, infosec, computer networks, cloud computing, and many other IT- and CS-related topics. Following graduation, he worked as a senior software engineer in Bangalore at SONY Indian Software Center, where he served as a full stack developer building and supporting web applications.
Since joining the IGB team, Kiran has contributed to many aspects of the code, notably IGB’s use of JavaFX and OSGi bundle loading. He also updated IGB’s use of a very old third-party library, which required finesse and deep knowledge of how the IGB project handles and manages library jar files that are not already OSGi-ready bundles.
As he gained skill, Kiran began writing IGB “case studies” slide decks, narratives explaining how he has tackled particularly interesting or difficult bug fixes or improvements.
Kiran now works as a Software Engineer II at Microsoft.
 Pranav Sanjay Tambvekar joined the lab in September, 2018 while studying Computer Science at UNC Charlotte. Prior to UNC Charlotte, he studied at Pune University, earning his Bachelor’s Degree in Engineering in 2014.
Pranav Sanjay Tambvekar joined the lab in September, 2018 while studying Computer Science at UNC Charlotte. Prior to UNC Charlotte, he studied at Pune University, earning his Bachelor’s Degree in Engineering in 2014.
Following graduation, he worked for three years as a Programmer/Analyst at Cognizant Technology Solutions, where he focused mainly on supporting applications in banking and finance. Some of his work there included developing on user-facing applications tracking trades-status (Angular) and developing REST APIs (Spring MVC, Jersey).
At UNC Charlotte, Pranav has further developed his interests in machine learning and data visualization, while continuing to build his skills as a full stack developer.
Pranav’s main contribution to the IGB project was developing the IGB App Store, using the Cytoscape App Store code base as a starting point.
Pranav now works as a Software Engineer for SAP. Check out his personal Web page (github.io) here.
 Sneha Ramesh Watharkar joined the Integrated Genome Browser development team in March 2018 during her second semester at UNC Charlotte.
Sneha Ramesh Watharkar joined the Integrated Genome Browser development team in March 2018 during her second semester at UNC Charlotte.
Sneha earned her Bachelor of Engineering and Computer Science and Engineering with Distinction from B V Bhoomaraddi College of Engineering and Technology in Hubli, Karnataka, India.
After earning her undergraduate degree, she worked for several years as a software engineer in Bangalore for such companies as Bharat Electronics Ltd, Practo Technologies, and Sears Holding of India. During her various positions, she worked with a range of technologies and tools we also use in the IGB project, such as Jira, Maven, and others.
Sheha played a key role in developing and planning the Integrated Genome Browser App Store. In addition to implementing functionality, she took the lead on planning or organizing development tasks. Sneha also contributed to javascript code running on BioViz.org that enables IGB to load data from external data sources such as Galaxy and the BioAnalytic resource developed by Nick Provart’s group at University of Toronto.
Sneha graduated from UNC Charlotte in December, 2018. After that, she joined Wayfair as a software developer, based on Boston.
 Deepti Joshi joined the Integrated Genome Browser team in Spring of 2017 during her first semester as a CS Masters student at UNC Charlotte.
Deepti Joshi joined the Integrated Genome Browser team in Spring of 2017 during her first semester as a CS Masters student at UNC Charlotte.
Deepti graduated from Pune University with a Bachelor of Engineering Degree in Information Technology. She then worked as a software engineer for two years at Tech Mahindra in India. Her work there focused on developing billing applications for AT&T and providing 24/7 support. In this role, she worked with technologies such as IBM Mainframes, COBOL, DB2, JCL, IMSDB.
Deepti received her MS in Computer Science in May 2018. She now works as a Java developer at Deutsche Bank in the Raleigh-Durham area in North Carolina.
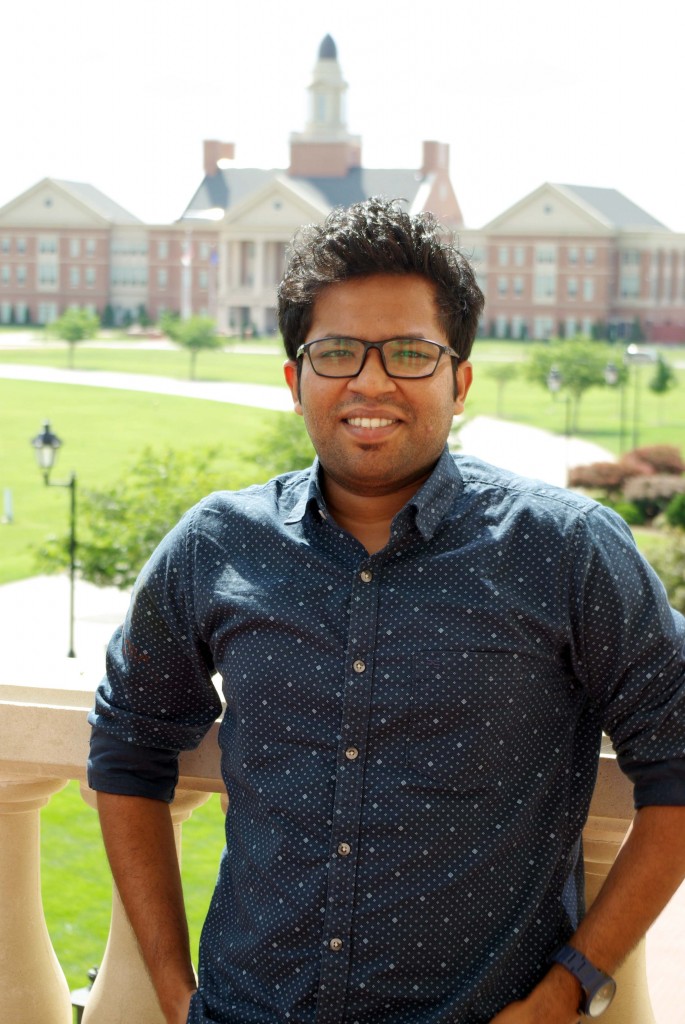 Sanket Patil joined the IGB development team in June of 2017 as a Graduate Research Assistant while working toward his Masters of Science degree in Information Technology. Before coming to Charlotte, he worked for two years as a software engineer at Prorigo Software Pvt. Ltd in India. Sanket holds a Bachelors degree in Instrumentation Technology from the University of Mumbai and a PG-Diploma in Advanced Computing from CDAC, also in Mumbai.
Sanket Patil joined the IGB development team in June of 2017 as a Graduate Research Assistant while working toward his Masters of Science degree in Information Technology. Before coming to Charlotte, he worked for two years as a software engineer at Prorigo Software Pvt. Ltd in India. Sanket holds a Bachelors degree in Instrumentation Technology from the University of Mumbai and a PG-Diploma in Advanced Computing from CDAC, also in Mumbai.
Since joining the project, Sanket made numerous contributions, everything from adding new functionality to improving our build and release process.
In addition to his work on IGB, Sanket contributed to CressExpress.org, a bioinformatics data mining site. His chief contribution was streamlining and updating the deployment process. Thanks to Sanket, CressExpress can now be easily deployed and tested using containers. In addition, Sanket deployed the CressExpress database on Amazon RDS, thus enabling “power users” to develop new data mining methods.
Sanket received his MS in Information Technology in May 2018. Since then, he has joined MicroStategy, based in the Washington DC area, as a software engineer.
Ashwini Kadam worked on the Integrated Genome Browser project from fall 2017 until fall of 2018.
She developed a prototype App for connecting to and reading BAM files from Google Drive called Google Connect. Ashwini also updated multiple out-dated user interface components in the IGB interface.
Following graduation, Ashwini moved to California, where she now works at Siemens PLM Software as a Software Engineer.
 Jennifer Daly joined the Lab as undergraduate student studying computer science and bioinformatics.
Jennifer Daly joined the Lab as undergraduate student studying computer science and bioinformatics.
During summer and fall of 2017, she worked as part of the Integrated Genome Browser team.
In addition to helping with code maintenance, she worked on building Apps to link IGB with cloud-based storage and computer resources such as Google Drive and CyVerse.
Jennifer now work as a Test Lead for Qlik.
 Devdatta Kulkarni joined the Integrated Genome Browser team in May 2016 to help develop IGB-fx while also maintaining IGB-Classic.
Devdatta Kulkarni joined the Integrated Genome Browser team in May 2016 to help develop IGB-fx while also maintaining IGB-Classic.
Devdatta is a graduate of Pune University, where he received a Bachelor’s degree in Engineering with a focus on Computer Engineering. As a student intern at Persistent Systems Ltd, he co-developed a DNA cryptography-based algorithm for secured data transfer over high speed networks. Following graduation, Devdatta re-joined the company as a software engineer. During his first studies at UNC Charlotte he has taken courses in visual analytics, big data analytics, cloud computing, and design.
Devdatta joined the technical staff at TIBCO Software, Inc after graduation. He now works as a software developer at Amazon.
David Norris received his Master’s degree in Computer Science from UNC Charlotte in 2012. Prior to this, he earned a Bachelor’s degree in History at UNC Charlotte. During his graduate career at UNC Charlotte, he added many new features to the Integrated Genome Browser, including several important improvements to the user interface. He also updated and modernized the PollenNetwork.org website.
Following graduation, he joined Red Hat as a software consultant. At Red Hat, he gained new expertise in linux server administration, JEE web application development, developing applications for cloud environments, cloud server administration, and agile software development methods. In February 2014, he rejoined the team as lead software developer for the Integrated Genome Browser project.
In early 2017, he left the lab to co-found Stackleader with fellow IGB project alumnus John Eckstein.
In his free time, David enjoys spending time with his family, running, and learning about new technologies and software tooling.
 Molly Crowder joined the group in summer of 2016 as a Graduate Research Assistant, focusing on analysis of alternative splicing.
Molly Crowder joined the group in summer of 2016 as a Graduate Research Assistant, focusing on analysis of alternative splicing.
Molly earned a BS in Biology from Wingate University in May of 2015. The following fall, she started the Professional Science Masters program in Bioinformatics at UNC Charlotte. Her first year in program, she took classes in data analysis, bioinformatics programming, databases, and genomic methods. Molly mastered these skills well enough to help Dr. Loraine teach Statistics for Bioinformatics, a required course in the Bioinformatics minor. She served as a Teaching Assistant for BINF 3121 Statistics for Bioinformatics in Fall of 2016 and Spring of 2017.
In Fall of 2017, Molly joined the Ph.D. program at University of Illinois, where she plans to student microbial ecology.
 Jessi Davis joined the group during the summer of 2016 as a student intern during her junior year at A.L. Brown High School in Kannapolis. Working closely with mentor Nowlan Freese, she investigated the function of alternatively spliced genes. She contributed to many other projects, as well, doing everything from tending plants to running gels.
Jessi Davis joined the group during the summer of 2016 as a student intern during her junior year at A.L. Brown High School in Kannapolis. Working closely with mentor Nowlan Freese, she investigated the function of alternatively spliced genes. She contributed to many other projects, as well, doing everything from tending plants to running gels.
She did great work, and we know she has a bright future in whatever field she decides to go into!
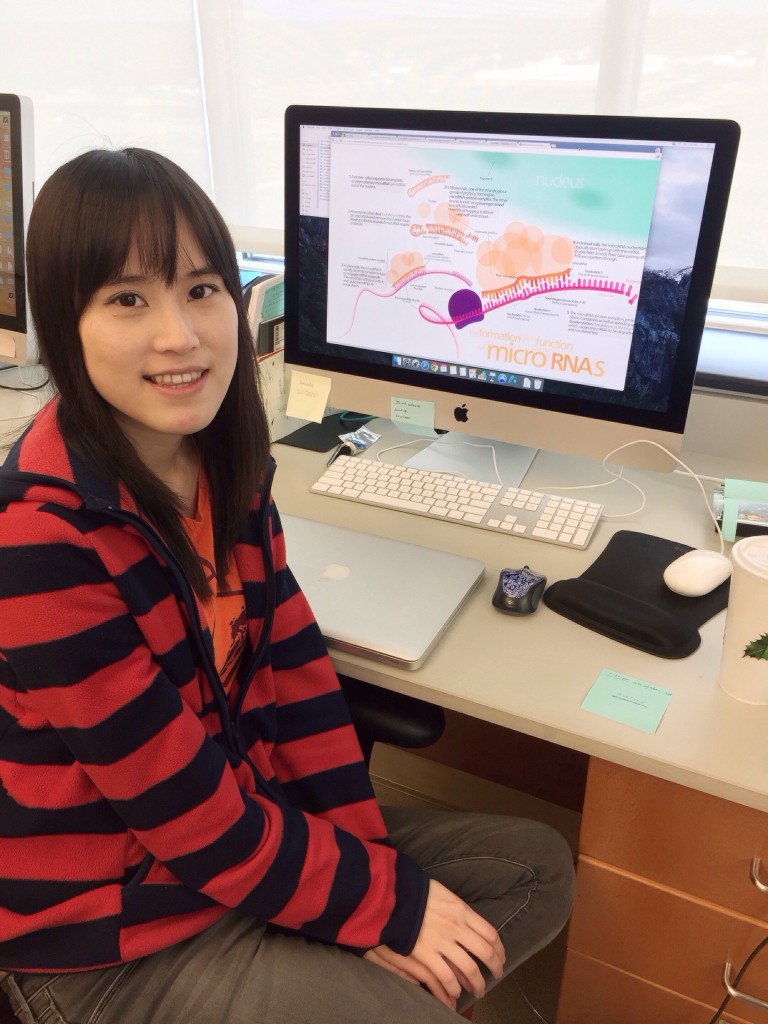 Ying-Chen Lin joined the lab in the fall semester of 2015. Working with April, she helped assess differential alternative splicing using fragment analysis on an ABI sequencer.
Ying-Chen Lin joined the lab in the fall semester of 2015. Working with April, she helped assess differential alternative splicing using fragment analysis on an ABI sequencer.
Prior to joining the lab, Ying-Chen earned a masters degree in horticulture from NC State University, working with Dr. Allan Brown, who has since moved to the International Institute of Tropical Agriculture in Arusha Tanzania.
Ying-Chen’s thesis research involved using our first blueberry genome releases (from May 2013) to develop and test new markers for a genetic map of blueberry. Working with the bioinformatics team, she then used her data to help us assess and validate a newer release of the genome assembly from August 2015.
Ying-Chen is now working on melon genetics as a Ph.D. student a Michigan State University.
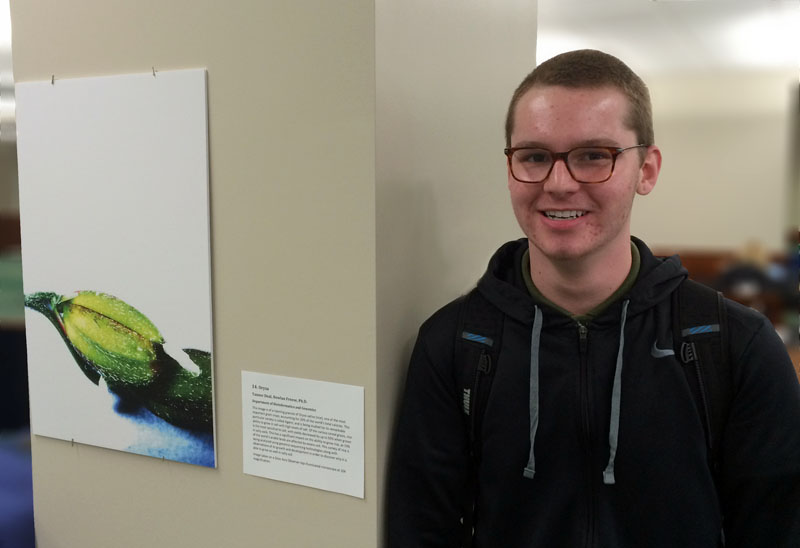 Tanner Deal joined the lab as a high school intern in the summer of 2015 and stayed on the following fall as a lab assistant. During his time in the lab, he helped with numerous projects, including our outreach activities introducing science to local students in the community.
Tanner Deal joined the lab as a high school intern in the summer of 2015 and stayed on the following fall as a lab assistant. During his time in the lab, he helped with numerous projects, including our outreach activities introducing science to local students in the community.
Tanner is now studying biology at UNC Charlotte.
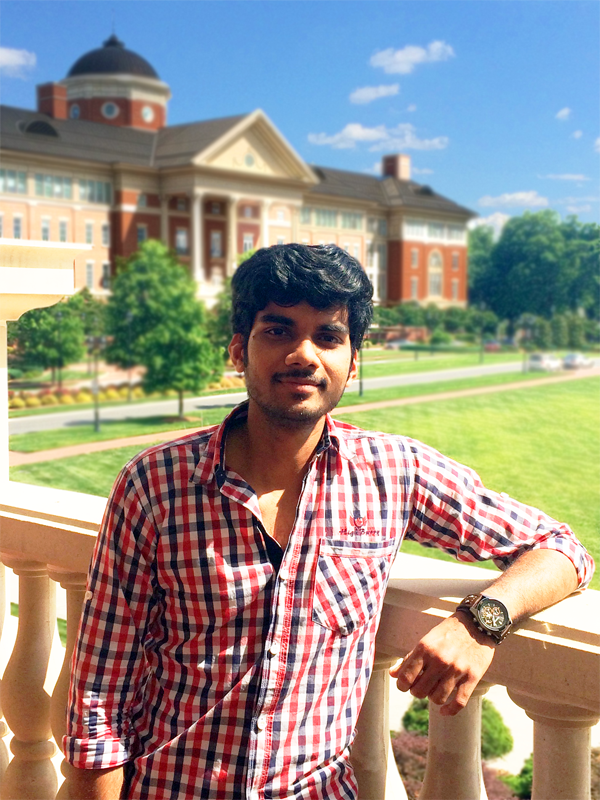 Muralidhar Akurati joined the Loraine Lab in May 2016, following his first semester in the CS Masters program at UNC Charlotte. Together with fellow CS student Devdatta Kulkarni, he helped to build and battle-test new versions of the IGB Apps platform – IGB-FX.
Muralidhar Akurati joined the Loraine Lab in May 2016, following his first semester in the CS Masters program at UNC Charlotte. Together with fellow CS student Devdatta Kulkarni, he helped to build and battle-test new versions of the IGB Apps platform – IGB-FX.
Muralidhar received his Bachelor of Engineering Degree from Andhra University in 2014. He then joined Tech Mahindra, a software development firm serving multiple clients worldwide. At Tech Mahindra, he worked on projects related to product integration, testing, and migrating software from .NET to Java.
Muralidhar now works as a Software Developer at Overstock.
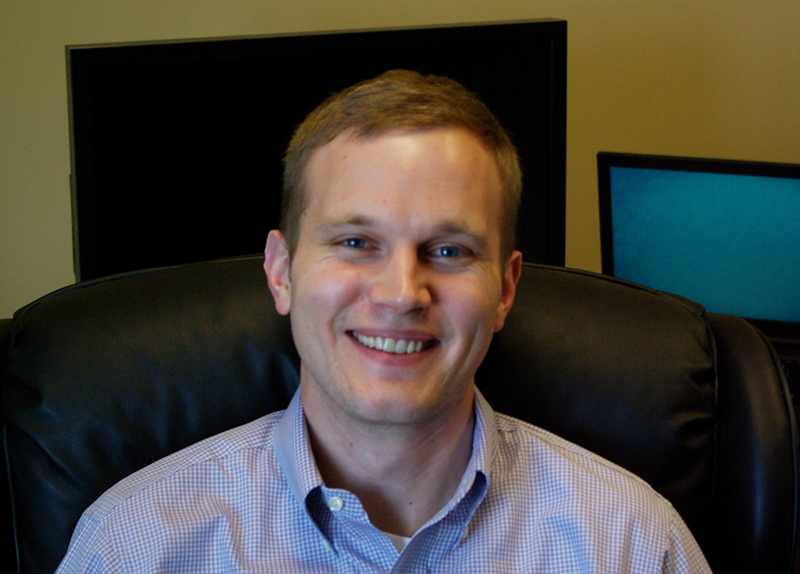 John Eckstein, a graduate of Virginia Tech University, worked for several years as a consultant for Red Hat, based in Atlanta. After learning about his former co-worker David Norris’ work on the IGB project, he decided to make the transition from consultant to full-time open source software developer here in the Loraine Lab. John contributed to several efforts, including re-vamping aspects of IGB’s services-based architecture, helping migrate IGB to JavaFX and building a new version of ProtAnnot that runs as an IGB plug-in.
John Eckstein, a graduate of Virginia Tech University, worked for several years as a consultant for Red Hat, based in Atlanta. After learning about his former co-worker David Norris’ work on the IGB project, he decided to make the transition from consultant to full-time open source software developer here in the Loraine Lab. John contributed to several efforts, including re-vamping aspects of IGB’s services-based architecture, helping migrate IGB to JavaFX and building a new version of ProtAnnot that runs as an IGB plug-in.
John now co-leads software consulting business Stackleader, based in Concord, NC.
 Daniella Triebwasser Freese worked with us on the IGB project during the spring semester of 2016 as a Bioinformatics Professional Science Masters student. Her first experience working on professional-grade software, she learned a huge amount and also contributed a huge amount. She helped us migrate the IGB code base to JavaFX and also entirely re-wrote the FindJunctions program, a core piece of our alternative splicing analysis infrastructure.
Daniella Triebwasser Freese worked with us on the IGB project during the spring semester of 2016 as a Bioinformatics Professional Science Masters student. Her first experience working on professional-grade software, she learned a huge amount and also contributed a huge amount. She helped us migrate the IGB code base to JavaFX and also entirely re-wrote the FindJunctions program, a core piece of our alternative splicing analysis infrastructure.
Danny Freese now works as a Software Engineering Consultant at Stackleader, co-founded by fellow IGB project alumni John Eckstein and David Norris.
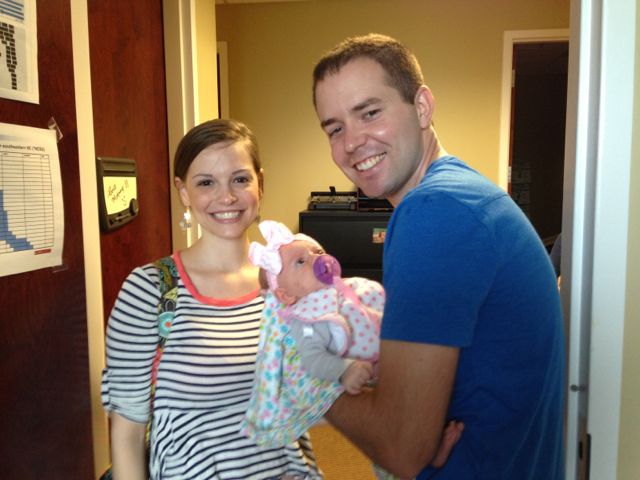
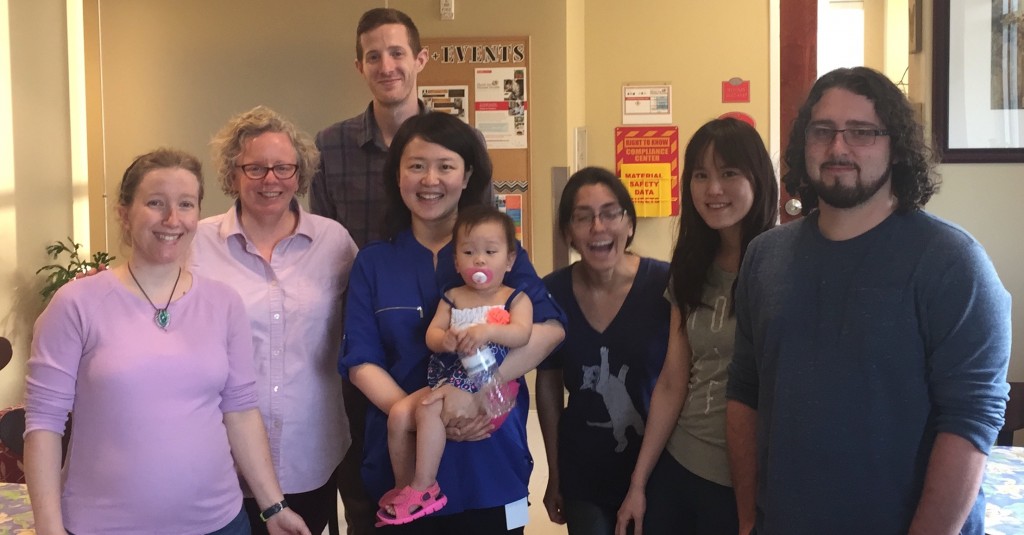 Jinjie Duan (center) from University of Aarhus in Denmark joined the lab as a visiting Ph.D. student in spring of 2016. In the photo, Jinjie is holding Amy, her one-year-old daughter.
Jinjie Duan (center) from University of Aarhus in Denmark joined the lab as a visiting Ph.D. student in spring of 2016. In the photo, Jinjie is holding Amy, her one-year-old daughter.
During her visit, Jinjie developed methods for improving gene annotations built from RNA-Seq data. She worked closely with Ann and Ivory (left) on a project investigating alternative splicing in rice. Before starting her PhD program, she worked at the Beijing Genomics Institute. At University of Aarhus, she has worked on genomics of python and spider.
Jinjie is now a bioinformatician at iPSYCH and the Department of Biomedicine at Aarhus University.
 Tarun Kumar Mall joined us in November, 2014. Tarun earned a Bachelors degree in Information Technology from Inderprastha Engineering College in June, 2011. Before arriving at UNC Charlotte in fall of 2014, he worked in commercial software development at L&T Infotech.
Tarun Kumar Mall joined us in November, 2014. Tarun earned a Bachelors degree in Information Technology from Inderprastha Engineering College in June, 2011. Before arriving at UNC Charlotte in fall of 2014, he worked in commercial software development at L&T Infotech.
While a programmer on the IGB project, Tarun implemented dozens of new features, contributed to the design of our new IGB services API, and wrote major parts of ProtAnnot, an IGB App for visualizing protein domains in the context of genomic sequence. During his tenure, he impressed us all with his love for programming, professionalism, generous spirit, and deep insight into all things code-related.
Tarun graduated in Dec. 2015 with a Masters in Information Technology at UNC Charlotte. He now works as a software engineer at Amazon.
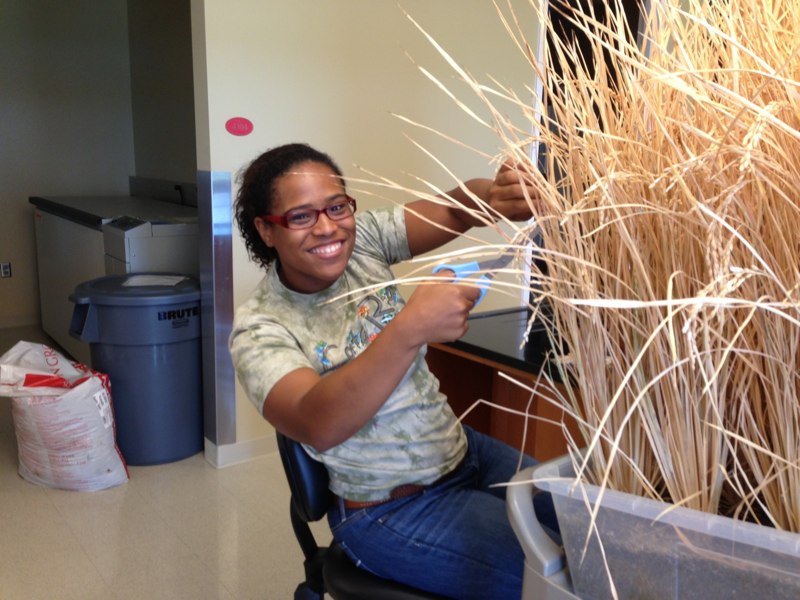 Britney David worked with us as a lab assistant while pursuing a degree in biology at UNC Charlotte. She also had many other jobs on campus, impressing us all with her energy and can-do spirit. In the Loraine lab, she helped with Arabidopsis crosses, rice harvesting, and lab maintenance tasks.
Britney David worked with us as a lab assistant while pursuing a degree in biology at UNC Charlotte. She also had many other jobs on campus, impressing us all with her energy and can-do spirit. In the Loraine lab, she helped with Arabidopsis crosses, rice harvesting, and lab maintenance tasks.
Britney is now pursuing career options in health sciences.
 Brock Overcash joined us in July of 2012 for a summer internship following his junior year of high school in Rowan County, which is north of the Research Campus.
Brock Overcash joined us in July of 2012 for a summer internship following his junior year of high school in Rowan County, which is north of the Research Campus.
During his internship, Brock helped out in the lab, improved the BioViz Web site, and did quality control analysis of RNA-Seq data.
After graduation, he attended college at Georgia Tech, majoring in computer science.
Brock is now a software engineer at Facebook.
 Hiral Vora started in the group as a masters student in Computer Science. One of his first projects was to re-write ProtAnnot, ultimately published as a new IGB plug-able App.
Hiral Vora started in the group as a masters student in Computer Science. One of his first projects was to re-write ProtAnnot, ultimately published as a new IGB plug-able App.
Following graduation, he rejoined the project as a staff software engineer, eventually taking over from John Nicol as the lead engineer on the IGB project.
During his tenure, he managed our collaboration with Genentech and helped mentor many students. His contributions to IGB and to biology are impossible to estimate.
Hiral is now a senior engineer at Deutsches Bank.
 Alyssa Gulledge, Ph.D. led outreach and testing efforts for the Integrated Genome Browser project from September 2010 to to December 2013. During her three year tenure, she took the lead on many critical efforts, including: updating and improving the IGB User’s Guide, designing new sequence alignment (BAM File) visualizations, developing better ways to test IGB in advance of new releases, and much more. She also helped us better understand users’ needs by constantly seeking their feedback and input on how they use IGB.
Alyssa Gulledge, Ph.D. led outreach and testing efforts for the Integrated Genome Browser project from September 2010 to to December 2013. During her three year tenure, she took the lead on many critical efforts, including: updating and improving the IGB User’s Guide, designing new sequence alignment (BAM File) visualizations, developing better ways to test IGB in advance of new releases, and much more. She also helped us better understand users’ needs by constantly seeking their feedback and input on how they use IGB.
In 2012 and 2013, she helped organize the UNC Charlotte Workshop in Next-Generation Sequencing. In this role, she managed development of the conference Web site, supervised and trained teaching assistants, coordinated speaker schedules, organized logistics, and co-ran the meeting.
She also found time to write articles on visualizing RNA-Seq data:
- A protocol for visual analysis of alternative splicing in RNA-Seq data using integrated genome browser. PubMed
- Mining Arabidopsis thaliana RNA-seq data with Integrated Genome Browser reveals stress-induced alternative splicing of the putative splicing regulator SR45a PubMed
Dr. Gulledge now works as a Grants and Contracts Writer at Kennesaw State University in Georgia.
Katy Kubiak worked with us during 2012 while earning her masters degree in Bioinformatics. She helped us as a Research Assistant and Software Tester, as well as TA for a workshop in Next-Generation Sequence Analysis.
Her main focus was on helping validate new releases of Integrated Genome Browser. She also contributed data analysis code and helped improve IGB training materials.
Katy now works as a Software Engineering Advisor at Cigna.
 Anuj Puram worked on the IGB project in 2012 while earning his masters degree in Computer Science at UNC Charlotte.
Anuj Puram worked on the IGB project in 2012 while earning his masters degree in Computer Science at UNC Charlotte.
Working closely with senior developer Hiral Vora, Anuj played a key role in developing the FindJunctions operator in IGB, which identifies and quantifies support for introns in RNA-Seq data. In addition, he developed the first version of the stand-alone FindJunctions tool, which we used for many years afterward.
Following graduation, Anuj has worked as a software consultant on a variety of projects.
Anuj is now an Ember JS Developer at Q2, in Austin, Texas.
 Praveen Babu Patchalla worked with us during 2012 while pursuing his Masters degree in Computer Science at UNC Charlotte. His main project was to maintain and add new features to the Pollennetwork.org website.
Praveen Babu Patchalla worked with us during 2012 while pursuing his Masters degree in Computer Science at UNC Charlotte. His main project was to maintain and add new features to the Pollennetwork.org website.
One such feature was an on-line search tool that enables project participants to add a listing of new pollen-related publications to the Web site. Praveen found a way to query a bibliographic database, thus saving us the effort of having to enter author, journal, date, and other article information by hand. He also helped improve the look of the site and helped us migrate it to a new Web server.
Following his graduation, Praveen joined the Janies lab at UNC Charlotte as a bioinformatics programmer, and then later accepted a position as a software developer at CNSI (Client Network Services, Inc.) in Maryland.
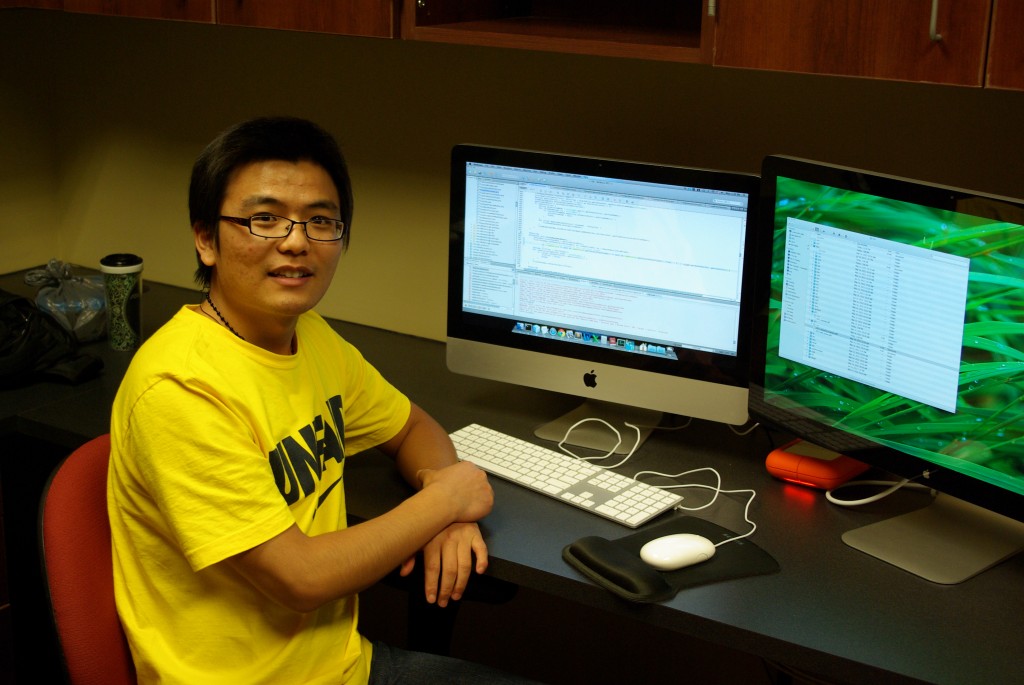 Fuquan Wang worked on the IGB project while pursuing his masters degree in Computer Science at UNC Charlotte. He added many features, notably the superimposition of amino acid sequences when users zoom in on protein-coding gene models. He added code that guarantees high contrast between amino acid letters and their background, a feature we’ve since replicated many times throughout IGB and related applications, like ProtAnnot.
Fuquan Wang worked on the IGB project while pursuing his masters degree in Computer Science at UNC Charlotte. He added many features, notably the superimposition of amino acid sequences when users zoom in on protein-coding gene models. He added code that guarantees high contrast between amino acid letters and their background, a feature we’ve since replicated many times throughout IGB and related applications, like ProtAnnot.
Following graduation, Fuquan moved to Detroit where he worked at Compuserve. He now works as a software development engineer at Amazon Web Services.
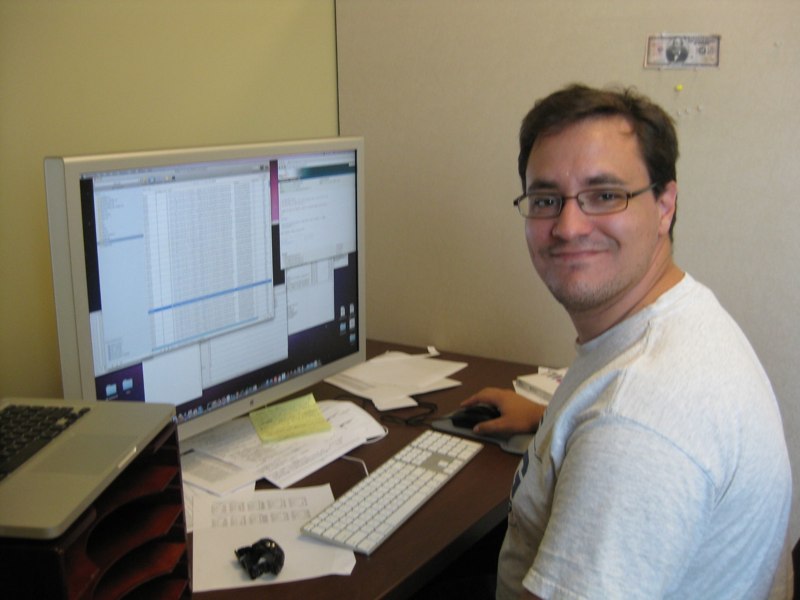 Adam Baxter joined us from 2011 to 2012 while working toward his Profesional Science Master degree in bioinformatics at UNC Charlotte. He contributed to the blueberry transcriptome project and was a Kannapolis Scholar, one of around 20 students who received a USDA fellowship to pursue graduate studies at the North Carolina Research Campus. In addition to doing research, Adam was active in the Bioinformatics Assembly of Students (BiAS) at UNC Charlotte and served as its fourth President.
Adam Baxter joined us from 2011 to 2012 while working toward his Profesional Science Master degree in bioinformatics at UNC Charlotte. He contributed to the blueberry transcriptome project and was a Kannapolis Scholar, one of around 20 students who received a USDA fellowship to pursue graduate studies at the North Carolina Research Campus. In addition to doing research, Adam was active in the Bioinformatics Assembly of Students (BiAS) at UNC Charlotte and served as its fourth President.
Following graduation, Adam joined Red Hat, where he works as a software consultant helping clients deploy open source infrastructure.
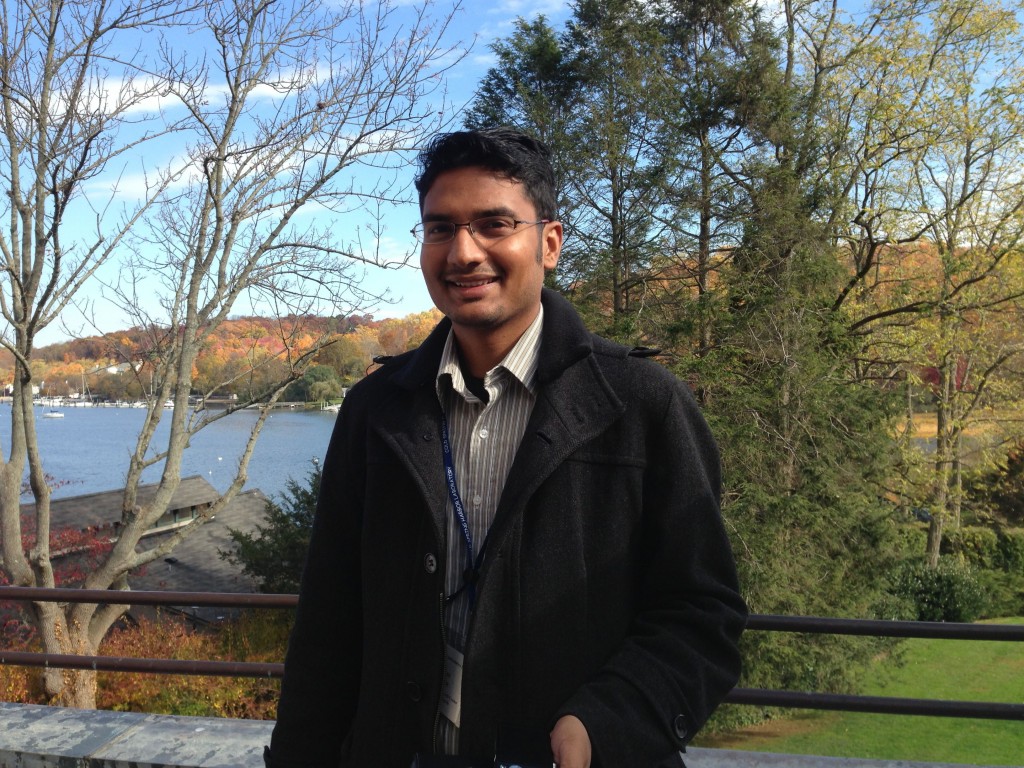 Vikas Gupta joined the lab from March to July 2013 as visiting student from University of Aarhus in Denmark. After earning his his Ph.D. degree in Bioinformatics and Computer Science in 2014, Vikas joined CLC Genomics, also in Denmark, as Bioinformatics Scientist.
Vikas Gupta joined the lab from March to July 2013 as visiting student from University of Aarhus in Denmark. After earning his his Ph.D. degree in Bioinformatics and Computer Science in 2014, Vikas joined CLC Genomics, also in Denmark, as Bioinformatics Scientist.
During his stay in the lab, Vikas worked on many projects, but his main contribution was to the blueberry transcriptome project. Already an expert on genome annotation thanks to his graduate studies, Vikas took the lead on using our RNA-Seq data to annotate the May 2013 draft of the blueberry genome.
Vikas Gupta is now works as a Bioinformatics Development Lead at Qiagen.
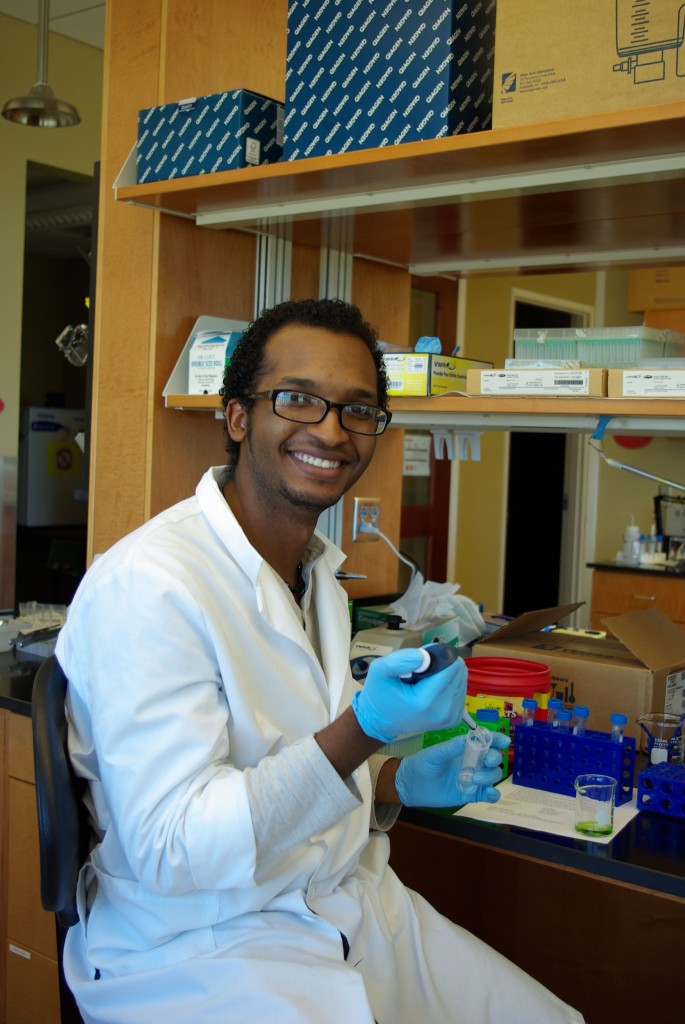 Darius Bost worked with us in 2012 during the summer of his sophomore year at NC A&T, where he studied mathematics and biology.
Darius Bost worked with us in 2012 during the summer of his sophomore year at NC A&T, where he studied mathematics and biology.
In addition to helping out with nearly every project then underway in the lab, Darius developed hands-on lab activities for high school students to learn about Mendelian genetics. One highlight of the summer was when Darius and the team hosted 30+ students and teachers visiting from Olympic High School in Charlotte. Using Arabidopsis floral mutants, participants learned about segregation ratios in genetics.
Darius Bost is now pursing his Ph.D. degree at UNC Chapel Hill.
 Vineeth Mylapur joined the group in 2012 as a graduate student in Computer Science at UNC Charlotte.
Vineeth Mylapur joined the group in 2012 as a graduate student in Computer Science at UNC Charlotte.
Vineeth continued development of the Pollen Research Coordination Network site PollenNetwork.org, taking over from Peter Pham.
He added several new features and helped improve the site’s utility as a community clearinghouse for pollen research information.
Following graduation, Vineeth joined Rackspace as an Oracle Business Analyst, based in San Antonio, Texas.
 Zhong (Nick) Ren worked with us from June 2011 until graduating in May 2012 with a Masters degree in Computer Science.
Zhong (Nick) Ren worked with us from June 2011 until graduating in May 2012 with a Masters degree in Computer Science.
A talented and dedicated developer, Nick improved many aspects of IGB. He improved the IGB bookmarking system, re-developed many facets of the IGB user interface, and also added a way for users to specify resolution of saved IGB images.
After graduation, Nick joined the Center for Human Genome Variation at Duke University, where he implemented bioinformatics data analysis pipelines.
Nick Ren is now a software engineer specializing in big data in bioinformatics. He works at Columbia University in New York.
 Nikhil Dahake worked on building a new version of CressExpress, a website devoted to mining Arabidopsis microarray data. This involved front-end web designing, back-end logic implementation, and database script generation.
Nikhil Dahake worked on building a new version of CressExpress, a website devoted to mining Arabidopsis microarray data. This involved front-end web designing, back-end logic implementation, and database script generation.
Nikhil Dahake now works as a Software Development Engineer II at TenMarks Education, an Amazon company.
 Tarun Santhosh Kanaparthi joined the Integrated Genome Browser team in fall of 2012, his first semester at UNC Charlotte. During his tenure, Tarun added many new features to the software, helped improve IGB usability, and helped test new versions prior to release. Tarun earned his MS in Computer Science in Dec. of 2014.
Tarun Santhosh Kanaparthi joined the Integrated Genome Browser team in fall of 2012, his first semester at UNC Charlotte. During his tenure, Tarun added many new features to the software, helped improve IGB usability, and helped test new versions prior to release. Tarun earned his MS in Computer Science in Dec. of 2014.
Following graduation, he accepted a position with Empirix, Inc in Boston. Empirix develops testing and monitoring tools for computer networks.
In addition to writing code, Tarun enjoys traveling and playing tennis.
Tarun Santosh Kanaparthi now works as a Software Engineer at Microsoft.
 Ivory Clabaugh Blakley worked with us from 2012 until 2018.
Ivory Clabaugh Blakley worked with us from 2012 until 2018.
A graduate of UNC Charlotte, she first joined the lab as a lab assistant, eventually earning a promotion to Research Technician and then Research Specialist.
At first, her role was mainly in the experimental biology lab; she helped run several experiments investigating how stresses affect splicing regulation in plants. Through taking classes and self-study, she gained new expertise in R programming and transitioned to data analyst.
During her tenure, she contributed to many articles, mainly working with collaborators Joe Kieber (UNC Chapel Hill) and Eric Schaller (Dartmouth) on regulation of cytokinin signaling in rice and Arabidopsis.
In her spare time, Ivory enjoys several hobbies, including bee-keeping, contra dancing, and tennis.
Ivory Blakley is now a Research Specialist in the Fodor Lab at UNC Charlotte.
 Roshanda Barner, Ph.D. joined us as a summer intern while studying math and biology at NC A&T University. During her first summer, she learned R and bioinformatics data analysis. She returned as a summer intern during the next three summers, eventually joining the Dept of Bioinformatics as a Ph.D. student.
Roshanda Barner, Ph.D. joined us as a summer intern while studying math and biology at NC A&T University. During her first summer, she learned R and bioinformatics data analysis. She returned as a summer intern during the next three summers, eventually joining the Dept of Bioinformatics as a Ph.D. student.
During her time in the Loraine Lab, Roshanda contributed to many projects, including co-expression analysis, RNA-Seq data analysis, and data wrangling for the IGB project.
Dr. Barner now is a Postdoctoral Scholar at University of Southern California at the Keck School of Medicine.
 Shira Stav, Ph.D. worked with us as an intern while taking classes and preparing for graduate school in biology. A physics major during her undergraduate studies, Shira became interested in biology and joined the Loraine Lab to get research experience in bioinformatics and biology. During her time in the lab, she worked with Smadar Harpaz-Saad
Shira Stav, Ph.D. worked with us as an intern while taking classes and preparing for graduate school in biology. A physics major during her undergraduate studies, Shira became interested in biology and joined the Loraine Lab to get research experience in bioinformatics and biology. During her time in the lab, she worked with Smadar Harpaz-Saad
(then a postdoc in Joe Kieber’s lab at UNC Chapel Hill) on co-expression analysis of Arabidopsis seed coat genes. She also helped out in the lab with collecting and processing samples for RNA-Seq, including Arabidopsis and blueberry.
After leaving the lab, Shira earned a Ph.D. degree from Yale University, where she developed sequence analysis methods to investigate RNA structure.
Dr. Stav has since returned to Charlotte, where she now works as a data scientist at Wells Fargo.
 Peter Pham earned a Professional Science Masters degree in in Bioinformatics in 2012. He worked with us during a summer internship, when he learned how to use DESeq to analyze RNA-Seq data. He also developed our first version of the PollenNetwork.org website. In addition to setting up the site, Peter experimented with adding custom modules to display and visualize RNA-Seq data.
Peter Pham earned a Professional Science Masters degree in in Bioinformatics in 2012. He worked with us during a summer internship, when he learned how to use DESeq to analyze RNA-Seq data. He also developed our first version of the PollenNetwork.org website. In addition to setting up the site, Peter experimented with adding custom modules to display and visualize RNA-Seq data.
Peter now works for a bioinformatics consulting company in Maryland.
 Afshan Jalali worked with us during the summer of 2011 while earning a masters degree in Bioinformatics at UNC Charlotte.
Afshan Jalali worked with us during the summer of 2011 while earning a masters degree in Bioinformatics at UNC Charlotte.
She developed the first version of the Pollen Research Coordination Network Web site at http://pollen.network.org, which uses the Drupal Content Management software. To develop the site, she worked closely with Pollen RCN members and served as a much-appreciated nexus to organize many people’s opinions and contributions.
Afshan is now a Senior Software Engineer at Illumina in San Diego.
 Vikram Bishnoi worked on the IGB project from 2011 to 2012 while earning his MS in Computer Science from UNC Charlotte. He made many contributions, including implementing the IGB Sequence Viewer, one of the unique features of the IGB software. The Sequence Viewer helps IGB users copy and translate genomic sequence data.
Vikram Bishnoi worked on the IGB project from 2011 to 2012 while earning his MS in Computer Science from UNC Charlotte. He made many contributions, including implementing the IGB Sequence Viewer, one of the unique features of the IGB software. The Sequence Viewer helps IGB users copy and translate genomic sequence data.
Vikram is now a Vice President at Bank of America Merrill Lynch, promoted from his original position as as Technology Analyst.
About the IGB project, Vikram says:
Developing code for IGB sharpened my skills on Core Java, Object oriented programming, and familiarized me with svn, and web, and application hosting. Also, it made me practice working in team, using several tools like JIRA for incident management, and visual analytics for effective representation of data. These skills cultivated and refined while working on IGB has helped me to expand my career opportunities and to add value to my role at my new job.
 Ehsan Tabari joined us as a rotation student during the summer of 2010.
Ehsan Tabari joined us as a rotation student during the summer of 2010.
During his rotation, he added many important new features to the IGB software. He also built a blueberry EST annotation Web site.
Ehsan then joined the laboratory of ZhengCheng Su on the main UNC Charlotte campus, where he is developed to methods to reduce effects of coverage bias in RNA-Seq data sets. He also continued to contribute ideas for improving IGB, such as a new type of coverage graph that plots the start position of read alignments.
Ehsan Tabari is now a Principal Scientist II at Roche Sequence Solutions.
 Nate Watson worked with us as a Graduate Student Researcher on the blueberry genome project while earning his Professional Science Masters degree in Bioinformatics.
Nate Watson worked with us as a Graduate Student Researcher on the blueberry genome project while earning his Professional Science Masters degree in Bioinformatics.
As an intern, he worked on on assembling 454 transcriptome data. He tested transcriptome assembly programs and developed a pipeline for quality screening and assembling the data. He also helped helped curate genome annotation and related data sets for the IGBQuickLoad.org data repository. As part of his work for the IGB project, he wrote code for converting GFF files to BED files.
Following graduation, Nate joined a plant biotechnology company in Davis, CA which later became part of Bayer Crop Sciences. Nate Watson now works as a Bioinformatician within Stanford Healthcare’s Clinical Genomics Program.
 Archana Raja was a graduate student researcher on the IGB project team while working toward her Professional Science Masters degree in Bioinformatics. A versatile and dedicated scientist, Archana contributed to many projects. She led our IGB testing efforts throughout her tenure, and in her spare time, led efforts to make and test a new “bpmap” file for an Arabidopsis tiling array from Affymetrix. Since graduation, she has worked on blueberry genomics for NC State, data analysis for University of Chicago, and human genetics at University of Washington in Evan Eichler’s lab.
Archana Raja was a graduate student researcher on the IGB project team while working toward her Professional Science Masters degree in Bioinformatics. A versatile and dedicated scientist, Archana contributed to many projects. She led our IGB testing efforts throughout her tenure, and in her spare time, led efforts to make and test a new “bpmap” file for an Arabidopsis tiling array from Affymetrix. Since graduation, she has worked on blueberry genomics for NC State, data analysis for University of Chicago, and human genetics at University of Washington in Evan Eichler’s lab.
She has also given guest lectures in the Bioinformatics Department’s Professional Development classes.
Archana Raja is now a Computational Biologist at Stanford University School of Medicine.
 Adam English joined the lab as a graduate student researcher while working toward his Professional Science Masters degree in Bioinformatics. He worked on alternative splicing in Arabidopsis and was the lead author of the ArabiTag software.
Adam English joined the lab as a graduate student researcher while working toward his Professional Science Masters degree in Bioinformatics. He worked on alternative splicing in Arabidopsis and was the lead author of the ArabiTag software.
Following graduation, Adam joined the Baylor Genome Center, where he now holds a position as Senior Bioinformatics Programmer. At Baylor, he wrote PBJelly, a popular tool improving genome assemblies using sequence data from the Pac Bio platform.
Adam English is now a Bioinformatics Scientist at Spiral Genetics, based in Seattle.
 John Nicol joined the team as lead software engineer on the Integrated Genome Browser Project.
John Nicol joined the team as lead software engineer on the Integrated Genome Browser Project.
During his tenure, John added dozens of features to IGB, re-factored the code base, introduced industry-standard software development practices, and helped mentor more than a dozen students, including IGB lead engineer Hiral Vora, who replaced John in 2011.
Since moving on in 2011, John has worked in software development at LinkedIn and Amazon. John Nicol is now a Staff Software Engineer at Duolingo.
 Kyle Suttlemyre worked with the IGB team to implement new ways for IGB to consume data via REST Web services.
Kyle Suttlemyre worked with the IGB team to implement new ways for IGB to consume data via REST Web services.
Inspired by collaborator Nick Schurch at University of Dundee, Kyle developed a simple but innovative technique using Javascript to send data to IGB from the wildly popular Galaxy workflow server.
Since then, we have extended this technique to other data sources, such as the Bio-Analytic Resource hosted at the University of Toronto.
Thank you Kyle for your great work! (You are a bioinformatics superhero.)
 April Roberts Estrada joined the group in 2011 as a Research Specialist to help manage the experimental side of the lab.
April Roberts Estrada joined the group in 2011 as a Research Specialist to help manage the experimental side of the lab.
During her tenure, she trained more than a dozen students and summer interns, brought new cloning and alternative splicing measurement techniques to the lab, and contributed to multiple software projects as a representative domain expert in biology.
She also published several articles, spoke about her work at national meetings, and helped run training workshops at UNC Charlotte and beyond.
Building on her experiences managing scientific data and documents, April joined Duke Energy in 2018 as a document control specialist.
 Ketan Patel, Ph.D. joined the lab in late 2008 as a postdoctoral researcher. He helped set up the laboratory and led our first sequencing experiments. Ketan played a key role in the blueberry genome project – he designed experiments, collected samples, made libraries, and led our first annotation efforts.
Ketan Patel, Ph.D. joined the lab in late 2008 as a postdoctoral researcher. He helped set up the laboratory and led our first sequencing experiments. Ketan played a key role in the blueberry genome project – he designed experiments, collected samples, made libraries, and led our first annotation efforts.
In 2011, he joined the US Naval Medical Research Center in Silver Spring, Maryland as a Research Scientist. During his tenure there, Dr. Patel and Navy colleagues set up and ran a mobile lab for diagnosing ebola infections in Africa. And in 2015, he was inducted into the Texas A&M University Academy of Distinguished Former Students.
Dr. Patel is now a Microbiologist at the Centers for Disease Control and Prevention in Atlanta.